COMP (CA): Szurmant Lab
Bacterial signal transduction systems and their role in physiology, antimicrobial resistance and utility as drug targets
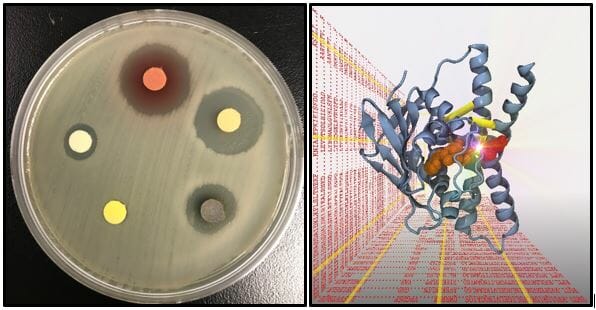
Pathogenic bacteria are some of the most formidable threats to human health. These threats appeared all but eliminated, thanks due to the discovery of powerful antibiotics. The constant exposure of bacteria to these antibiotics has selected for potent multi-drug resistant bacteria, so-called superbugs that are making a strong comeback. To cope with this renewed threat a dedicated effort by the scientific community is needed to identify new drug targets and to generate inhibitors of such targets. The Szurmant laboratory contributes to this endeavor by studying essential aspects of bacterial physiology and signal transduction in model bacteria and selected pathogens. A unique feature of the lab is the integration of information stemming from numerous disciplines, including structural biology, genetics, molecular biology, bioinformatics and biophysics.
Bacterial signal transduction and its role in important physiological and developmental processes
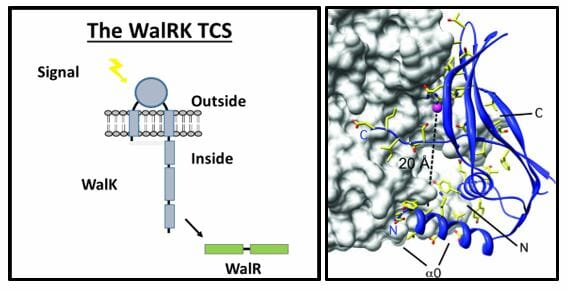
The Szurmant laboratory has a long-standing interest in bacterial signal transduction. The focus is on the so-called two-component system, a mechanism that is widely utilized by bacteria in many developmental decisions, such as sporulation, genetic competence development, chemotaxis. From a medical perspective, those systems that contribute to antimicrobial resistance, virulence factor expression and those essential for viability are of particular interest. The laboratory has a special focus on the study of WalRK, the only essential two-component signal transduction system of bacteria in the phylum Firmicutes. This phylum includes significant human pathogens such as Staphylococcus aureus, Streptococcus pyogenes, Enterococcus faecalis and bioterrorism thread Bacillus anthracis. Using molecular genetics techniques, we have identified the role this system plays in bacterial physiology. Some efforts in the lab are directed to WalRK’s involvement in antimicrobial resistance of clinical isolates and testing its utility as an antimicrobial drug target.
Molecular evolution and protein-protein interactions
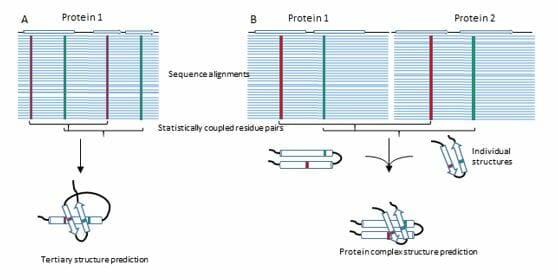
Proteins in all organisms undergo concerted interactions in order to execute catalytic, structural, transport and many other essential functions. Numerous genetic diseases are caused by mutations that interfere with wild type protein-protein interactions (PPI). Knowledge on all viral-host cell PPIs, such as those important for COVID-19 infection and disease, would greatly speed up new drug discovery. In bacteria, knowledge on all essential PPIs would greatly enhance the complement of antimicrobial drug targets, in the light of multi-drug resistant bacteria. We developed a widely applicable technology termed Direct Coupling Analysis (DCA), which in combination with experimental techniques has found wide application in the identification of PPI partners and in solving structures of proteins, protein complexes and alternative conformations of signaling proteins. Direct Coupling Analysis relies on the exponentially growing protein sequence databases to mathematically extract contact residue information within and between proteins. We are currently moving this technology forward by trying to understand the amino acid sequence code that dictates not only the structure of PPIs, but also their strength. To this end, we are experimentally sampling directed mutant libraries of thousands of variants of essential interacting protein pairs. In vivo evolution of libraries will inform about the epistatic coupling of amino acid residues on the contact surface between these proteins. Paired with DCA-based computational efforts this approach will provide a leap in our understanding of the protein-sequence code that dictates the strength of an interaction to one day predict all elements of protein function, relying solely on protein sequence. These efforts hold the long-term promise to help identify genetic determinants of complex diseases based on rare variants. In the context of antibiotic resistance, it can be helpful for predicting patterns of adaptive mutations of pathogens (bacteria and viruses) and contribute to the discovery of new therapeutic strategies.
Kim, I.M. and Szurmant, H. A Bacterial Goldilocks Mechanism. Elife. 24: e54244, 2020.
Rosales-Hurtado, M., Meffre, P., Szurmant, H., Benfodda, Z. Synthesis of Histidine Kinase Inhibitors and Their Biological Properties. Med. Res. Rev. 40: 1440-1495, 2020.
Croce, G., Gueudré, T., Ruiz Cuevas, M.V., Keidel, V., Figliuzzi, M., Szurmant, H. and Weigt, M. A Multi-Scale Coevolutionary Approach to Predict Interactions between Protein Domains. PLoS Comput. Biol. 15: e1006891, 2019.
Szurmant, H. Evolutionary Couplings of Amino Acid Residues Reveal Structure and Function of Bacterial Signaling Proteins. Mol. Microbiol. 112: 432-437, 2019.
Szurmant, H. and Weigt, M. Inter-Residue, Inter-Protein and Inter-Family Coevolution: Bridging the Scales. Curr. Opin. Struct. Biol. 2018:26-32, 2018.
Uguzzoni, G., John Lovis, S., Oteri, F., Schug, A.*, Szurmant, H.* and Weigt, M.* Large-Scale Identification of Coevolution Signals across Homo-Oligomeric Protein Interfaces by Direct Coupling Analysis. Proc. Natl. Acad. Sci. USA. 114: E2662-E2671, 2017. *Co-corresponding authors.
Boibessot, T., Zschiedrich, C.P., Lebeau, A., Bénimèlis, D., Dunyach-Rémy, C., Lavigne, J.P., Szurmant, H.*, Benfodda, Z.* and Meffre, P.*. The Rational Design, Synthesis, and Antimicrobial Properties of Thiophene Derivatives That Inhibit Bacterial Histidine Kinases. J. Med. Chem. 59: 8830-8847, 2016. *Co-corresponding authors
Zschiedrich, C. P., Keidel, V. and Szurmant, H. Molecular Mechanisms of Two-component Signal Transduction. J. Mol. Biol. 428: 3752-75, 2016.
Feinauer, C., Szurmant, H., Weigt, M. and Pagnani, A. Inter-Protein Sequence Co-Evolution Predicts Known Physical Interactions in Bacterial Ribosomes and the Trp Operon PLoS ONE. 11: e0149166, 2016.
Pokkuluri P.R., Dwulit-Smith, J., Duke, N.E., Wilton, R., Mack, J.C., Bearden, J., Rakowski, E., Babnigg, G., Szurmant, H., Joachimiak, A., and Schiffer, M. Analysis of Periplasmic Sensor Domains from Anaeromyxobacter dehalogenans 2CP-C: Structure of One Sensor Domain from a Histidine Kinase and Another from a Chemotaxis Protein. Microbiology Open. 2: 766-77, 2013.
Wu, R., Gu, M., Wilton, R., Babnigg, G., Kim, Y., Pokkuluri, P.R., Szurmant, H., Joachimiak, A., Schiffer, M. Insight into the Sporulation Phosphorelay: Crystal Structure of the Sensor Domain of Bacillus subtilis Histidine Kinase KinD. Protein Sci. 22: 564-76, 2013.
Szurmant, H. and Hoch, J.A. Statistical Analyses of Protein Databases Identify Structures and Mechanisms in Signal Activation of Sensor Histidine Kinases. Mol. Microbiol. 87: 707-12, 2013.
Dago, A.E., Schug, A., Procaccini, A., Hoch, J.A., Weigt, M., and Szurmant, H. The Structural Basis of Histidine Kinase Autophosphorylation: Integrating Genomics, Molecular Dynamics and Mutagenesis. Proc. Natl. Acad. Sci USA 109: E1733-42, 2012.
Diaz, A.R., Core, L., Jiang, M., Morelli, M., Chiang, C.H., Szurmant, H., and Perego, M. Bacillus subtilis RapA Phosphatase Domain Interaction with its Substrate, Phosphorylate Spo0F, and its Inhibitor, the PhrA peptide. J. Bacteriol. 194: 1378-88, 2012.
Fukushima, T., Furihata, I., Emmins, R., Daniel, R.A., Hoch, J.A., and Szurmant, H. A Role for the Essential YycG Sensor Histidine Kinase In Sensing Cell Division. Mol. Microbiol. 79: 503-22, 2011.
Procaccini, A., Lunt, B., Szurmant, H., Hwa, T., and Weigt, M. Dissecting the Specificity of Protein-Protein Interaction in Bacterial Two-Component Signaling: Orphans and Crosstalks. PLoS One 6: e19729, 2011.
Wilson, A.C. and Szurmant, H. Transposon Systems for Random Mutagenesis of Bacillus subtilis. Methods Mol. Biol. 765: 359-71, 2011.
Szurmant, H. and Hoch, J.A. Interaction Fidelity in Two-Component Signaling. Curr. Opin. Microbiol. 13: 190-7, 2010.
Schug, A., Weigt, M., Hoch, J.A., Onuchic, J.N., Hwa, T., and Szurmant, H. Computational Modeling of Phosphotransfer Complexes in Two-Component Signaling. Methods Enzymol. 471: 43-58, 2010.
Lunt, B., Szurmant, H., Procaccini, A., Hoch, J.A., Hwa, T., Weigt, M. Inference of Direct Residue Contacts in Two-Component Signaling. Methods Enzymol. 471: 17-41, 2010.
Chang, C., Tesar, C., Gu, M., Babnig, G., Joachimiak, A., Pokkuluri, P.R., Szurmant, H., and Schiffer, M. Periplasmic PAS Domains: A Dominant Structural Family Governing Signal Transduction. J. Bacteriol. 192: 1156-9, 2010.
Schug, A., Weigt, M., Onuchic, J.N., Hwa, T., and Szurmant, H. High Resolution Protein Complexes From Integrating Genomic Information With Molecular Simulation. Proc. Natl. Acad. Sci. USA 106: 22124-9, 2009.
Weigt, M., White, R. A., Szurmant, H., Hoch, J.A., and Hwa, T. Identification of Direct Residue Contacts in Protein-Protein Interaction By Message Passing. Proc. Natl. Acad. Sci. USA 106: 67-72, 2009.
Szurmant, H., Bobay, B.G., White, R. A., Sullivan, D.M., Thompson, R.J., Hwa, T., Hoch, J.A., and Cavanagh, J. Co-evolving Motions at Protein-Protein Interfaces of Two- Component Signaling Systems Identified by Covariance Analysis. Biochemistry, 47: 7782-4, 2008. Selected as “hot article” by the editorial board.
Fukushima, T., Szurmant, H., Kim, E.J., Perego, M., and Hoch, J.A. A Sensor Histidine Kinase Coordinates Cell Wall Architecture with Cell Division in Bacillus subtilis. Mol. Microbiol. 69: 621-32 (Including Cover Image), 2008.
Szurmant, H., Bu, L., Brooks, C.L., and Hoch, J.A. An Essential Sensor Histidine Kinase Controlled By Transmembrane Helix Interaction with its Auxiliary Proteins. Proc. Natl. Acad. Sci. USA 105: 5891-6, 2008.
Szurmant, H., White, R.A., and Hoch, J.A. Sensor Complexes Regulating Twocomponent Signal Transduction. Curr. Opin. Struct. Biol. 17: 706-15, 2007.
Santelli, E., Liddington, R.C., Mohan, M.A., Hoch, J.A., and Szurmant, H. The Crystal Structure of Bacillus subtilis YycI Reveals a Common Fold for Two Members of an Unusual Class of Histidine Kinase Regulatory Proteins. J. Bacteriol. 289: 3280-9, 2007.
Szurmant, H., Mohan, M.A., Imus, P.M., and Hoch, J.A. YycH and YycI Interact to Regulate the Essential YycFG Two-Component System. J. Bacteriol. 289: 3290-5, 2007.
White, R. A., Szurmant, H., Hoch, J.A., and Hwa, T. Features of Protein-protein Interactions in Two-Component Signaling Deduced from Genomic Libraries. Methods Enzymol. 422: 75-101, 2007.
Szurmant, H., Fukushima, T., M.A., and Hoch, J.A. The Essential YycFG Two- Component System in Bacillus subtilis. Methods Enzymol. 422: 396-417, 2007.
Szurmant, H., Zhao, H., Mohan, M.A., Hoch, J.A., and Varughese, K.I. The Crystal Structure of YycH Involved in the Regulation of the Essential YycFG Two-Component System in Bacillus subtilis reveals a Novel Tertiary Structure. Protein Sci. 15: 929-34 (Including Cover Image), 2006.
Wörner, K., Szurmant, H., Chiang, C., and Hoch, J.A. Phosphorylation and Functional Analysis of the Sporulation Initiation Factor Spo0A from Clostridium botulinum. Mol. Microbiol., 59: 1000-12, 2006.
Szurmant, H., Nelson, K., Kim, E.J., Perego, M., and Hoch, J.A. YycH Regulates the Activity of the Essential YycFG Two-Component System in Bacillus subtilis. J. Bacteriol. 187: 5419-26, 2005.
Szurmant, H. and Ordal, G.W. The Diversity in Chemotaxis Mechanisms among the Bacteria and Archaea. Microbiol. Mol. Biol. Rev. 68: 301-19, 2004.
Szurmant, H., Muff, T., and Ordal, G.W. Bacillus subtilis CheC and FliY are Members of a Novel Class of CheY-P Hydrolyzing Proteins in the Chemotactic Signal Transduction Cascade. J. Biol. Chem. 279: 21787-92, 2004.
Szurmant, H., Bunn, M.W., Cho, S.H., and Ordal, G.W. Ligand Induced Conformational Changes in the Bacillus subtilis Chemotaxis Receptor McpB Determined by Disulfide Crosslinking in vivo. J. Mol. Biol. 344: 919-28, 2004.
Szurmant, H., Bunn, M.W., Cannistraro, V.J., and Ordal, G.W. Bacillus subtilis Hydrolyzes CheY-P at the Place of its Action, the Flagellar Switch. J. Biol. Chem. 278: 48611-6, 2003.
Zimmer, M.A., Szurmant, H., Saulmon, M.M., Collins, M. A., Bant, J.S., and Ordal, G.W. The Role of Heterologous Receptors in McpB-Mediated Signaling in Bacillus subtilis Chemotaxis. Mol. Microbiol. 45: 555-69, 2002.


